[page edited on August, 24th, 2020]
Mapsembler2 is not maintained anymore
Sadly, we have no more man power to maintain and to update Mapsembler2. The last (2014) version is this one: mapsembler2_2.2.4.
Many bug fixes, new features, and optimizations are expected. Any help would be warmly welcome!
Mapsembler2 description and possible usages:
Mapsembler2 is a targeted assembly software. It takes as input any number of NGS raw read set(s) (fasta or fastq, gzipped or not) and a set of input sequences (starters). For each starter, Mapsembler2 outputs its sequence neighborhood as a linear sequence or as a graph, depending on the user choice. Mapsembler2 may be used for (not limited to): · Validate an assembled sequence (input as starter), e.g. from a de Bruijn graph assembly where read-coherence was not enforced. Checks if a known enzyme is present in a metagenomic NGS read set. · Enrich unmappable reads by extending them, possibly making them mappable · Checks what happens at the extremities of a contig · Check the presence / absence and quantify RNA seq splicing events. Check the presence/absence of SNPs or structural variants, …
What’s new in Mapsembler2?
Two main novelties are implemented in mapsembler2:
- The Minia data structure is now used for indexing reads. For the price of using a few Gigabytes RAM, Mapsembler2 is much faster as it reads the read files only once.
- The starters are only extended. Mapsembler does not search for all (possibly numerous) sub-starters.
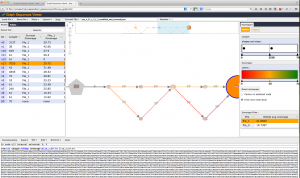
- An ad hoc visualization interface is proposed.
Video Tutos
Mapsembler2 install and use: http://www.youtube.com/watch?v=136mr5TOPCw
Mapsembler2 Graph of Sequence Viewer: http://www.youtube.com/watch?v=IEsE-iMNTJo
Download Mapsembler2:
Please read and accept the GNU Affero General Public License before use and diffusion. Any feedback and comment is warmly encouraged.
Actual version: mapsembler2_2.2.4
Logs:
- mapsembler2_2.2.4: fixes k-mer counting problems. Use by default 4GB memory
- mapsembler2_2.2.3: fixes compilation problems (tr1/ zlib / clang / boost)
- mapsembler2_2.2.2: Add compatibility with old version of gcc (min version :4.4.0) – Add the possibility to save/restore comments and highlights on GSV – Fix several minor bug
- mapsembler2_2.2.1: Cosmetic documentation and pipeline modifications – Reduced the archive size.
- mapsembler2_2.2.0.zip: Fixes a few bugs – improved the documentation –
- mapsembler2_2.1.5: Add options to set type of process, maximum graph depth and maximum number of nodes. The visualisation is now more simple and more stable.
- mapsembler2_2.1.2: much work has been done ! The visualization interface is now in beta test. The full package is operational.
- 2.0.5: mapsembler2_2.0.5. fixes a few bugs – A newer version integration new features and results visualization will be available in January.
- 2.0.4: mapsembler2_2.0.4 add the possibility to compile for larger k values, e.g. make k=55 – remove compilation bug.
- 2.0.3: mapsembler2_2.0.3: Added option -m, stops earlier if no substarter is generated
- 2.0.2 – mapsembler2_20130516: Correction of a bug in the sequencing error detection / correction
- 2.0.1 – mapsembler2_20130415: fixed a memory bug (thanks to Peter Foster).
- 2.0.0 – Initial version: mapsembler2_20130509 (beta version)
Download Mapsembler1 – from old web site: Alcovna
Please read and accept the License before use and diffusion. Any feedback and comment is warmly encouraged. The last mapsembler1 release may be downloaded here: mapsembler_1.3.21 (beta version).
Download Graph of Sequence Viewer for Galaxy:
GSV for Galaxy v1.0
GSV for Galaxy (beta version)
Contact
For any questions, contact us: pierre.peterlongo@inria.fr or on Biostars
